Mechanistic toxicology
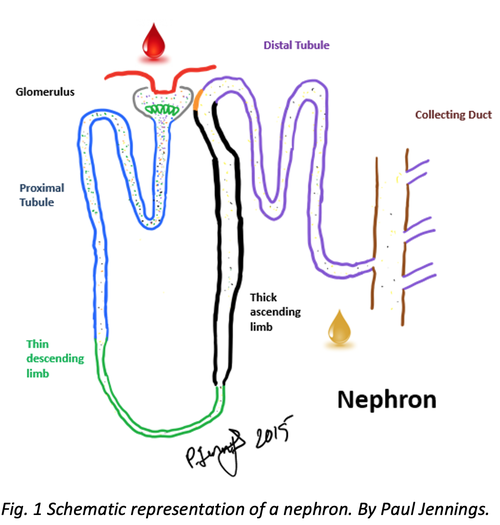
Nephrotoxicity assessment and mechanistic toxicology are the main specialty of the Molecular and Computational Toxicology research group at VU Amsterdam (Fig. 1). The laboratory is specialized in human cell culture, renal cell culture and iPSC derived cell culture coupled with morphological evaluation (high content imaging), electrical parameters (xCELLigence and transepithelial electrical resistance), mitochondrial respiration (Seahorse bioanalyser), toxicogenomics, biokinetics, stress response evaluation, and cyto-toxicity evaluation.
We provide specialist expert-based services in nephrotoxicity assessment (mechanistic toxicology) including human cell culture, renal cell culture and iPSC derived cell culture (Fig. 2 and Fig. 3) coupled with morphological evaluation (high content imaging, Fig. 4), electrical parameters (xCELLigence and transepithelial electrical resistance), mitochondrial respiration (Seahorse bioanalyser), toxicogenomics, biokinetics, stress response evaluation, and cyto-toxicity evaluation.
Research service areas include:
- Phase I and phase II metabolism and transport (phase III), including the influences of polymorphisms
- Molecular mechanisms of chemical- and nanomaterial-induced cellular perturbations
- Computational methods for metabolism prediction, site of metabolism and docking
- Mechanism of compound induced nephrotoxicity from repeat dose in vitro exposures utilizing integrated omics. (Wilmes et al. 2013b, 2015)
- Mitochondrial toxicity evaluation of agrochemicals. (van der Stel et al. 2020a)
- Renal transport studies. (Aschauer et al. 2015a)
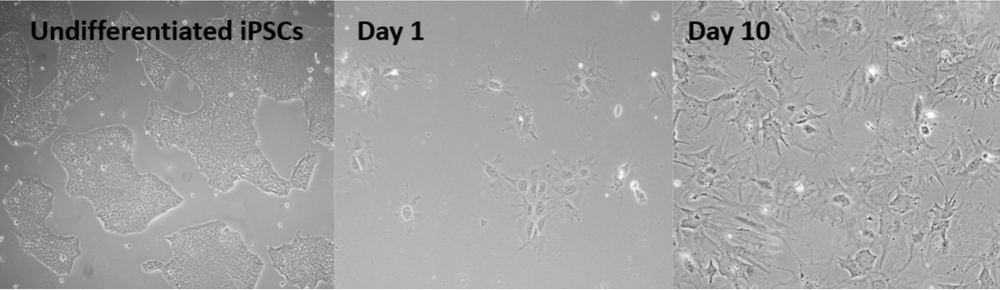
Fig.2 iPSC derived podocytes on differentiation day 1 and 10 (pictures provided by Cormac Murphy, c.murphy@vu.nl)
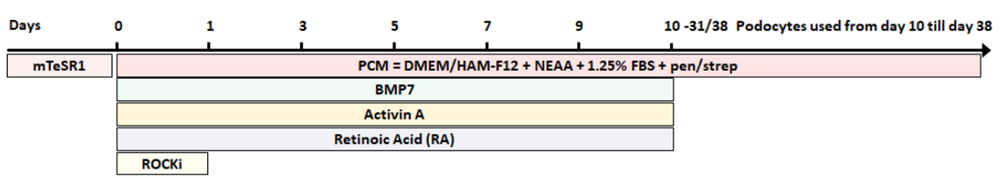
Fig. 3 Schematic representation of the VU podocyte differentiation protocol by Cormac Murphy. c.murphy@vu.nl
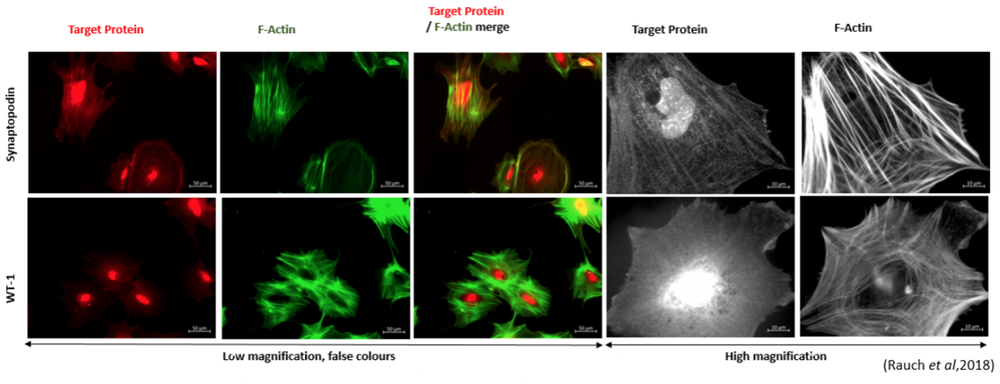
Fig. 4 Immunostaining of podocyte-like cells derived from the iPSC line SBAD2 with podocyte markers synaptopodin, WT-1 and F-actin staining (from Rauch et al, 2018).
Assays
Lactate. Alterations in supernatant lactate production correlates to glycolysis rates. Enhanced glycolysis is an excellent stress marker in cells with high rates of oxidative metabolism. It is measured in the collected supernatant via a colorimetric (in house assay) using any appropriate plate reader (Limonciel et al. 2011).
Resazurin reduction. Viability assay based on the ability of living cells to convert resazurin to the highly fluorescent product resorufin (Koppelstaetter et al. 2004).
Mitochondrial respiration. Label free oxygen consumption rate (OCR) and medium acidification (ECAR) in living cells (Seahorse XF bioanalyser) (van der Stel et al. 2020b).
High content imaging (HCI). Label free (digital phase contrast, bright field and fluorescent reporter-based assays), live staining and fixed cell imaging with confocal possibilities and advanced analysis algorithms (Operetta, PerkinElmer).
Cellular Impedance. The xCELLigence apparatus measures net impedance of each well, reported as Cell Index (CI), with high temporal resolution CI dependent on the amount of cells attached to the well, the forces at which they bind and the net volume of the cells per well and is considered a sensitive label-free viability assay (Limonciel et al. 2018).
Toxicogenomics. Quantification of thousands of individual mRNA molecules in human cells (Jennings et al. 2013; Aschauer et al. 2015b).
Biokinetics. Quantification of applied parent chemical and/or their metabolites in the cell culture supernatant, plastic and cells over time (HPLC, LC/MS-TOF, ICP-MS) (Wilmes et al. 2013a, 2015; Bellwon et al. 2015).
VISIT OUR SHOP
Request an Assay from Molecular and Computational Toxicology department of VU Amsterdam
Murphy C., Feifel E., Jennings P., Gstraunthaler G., Wilmes A. (2019) A Protocol for One-Step Differentiation of Human Induced Pluripotent Stem Cells into Mature Podocytes. In: Mandenius CF., Ross J. (eds) Cell-Based Assays Using iPSCs for Drug Development and Testing. Methods in Molecular Biology, vol 1994. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9477-9_8
Rauch C, Feifel E, Kern G, Murphy C, Meier F, Parson W, et al. (2018) Differentiation of human iPSCs into functional podocytes. PLoS ONE 13(9): e0203869. https://doi.org/10.1371/journal.pone.0203869
Wieser M, Stadler G, Jennings P, Streubel B, Pfaller W, Ambros P, Riedl C, Katinger H, Grillari J, Grillari-Voglauer R. hTERT alone immortalizes epithelial cells of renal proximal tubules without changing their functional characteristics. Am J Physiol Renal Physiol. 2008 Nov;295(5):F1365-75. doi: 10.1152/ajprenal.90405.2008. Epub 2008 Aug 20. PMID: 18715936.
Wilmes A, Bielow C, Ranninger C, et al (2015) Mechanism of cisplatin proximal tubule toxicity revealed by integrating transcriptomics, proteomics, metabolomics and biokinetics. Toxicol Vitr 30:117–127. https://doi.org/10.1016/j.tiv.2014.10.006
Wilmes A, Limonciel A, Aschauer L, et al (2013) Application of integrated transcriptomic, proteomic and metabolomic profiling for the delineation of mechanisms of drug induced cell stress. J Proteomics 79:180–194. https://doi.org/10.1016/j.jprot.2012.11.022